 Discard the flow through.Īdd 800 µl of ZymoPURE™ Wash 2 to the Zymo-Spin™ II-P Column and centrifuge at 6,000 rpm (~5,900 x g) for 1 min. Discard the flow through.Īdd 800 µl of ZymoPURE™ Wash 1 to the Zymo-Spin™ II-P Column and centrifuge at 6,000 rpm (~5,900 x g) for 1 min. Incubate the Zymo-Spin™ II-P/Collection Tube assembly at room temperature for 2 minutes and then centrifuge at 6,000 rpm (~5,900 x g) for 1 min. Place a Zymo-Spin™ II-P Column in a Collection Tube and transfer the entire mixture from step 8 into the Zymo-Spin™ II-P Column. Mix thoroughly by inverting the capped tube 8 times. Transfer 600 µl of supernatant from step 6 into a clean 1.5 ml microcentrifuge tube.īe careful not to disturb the yellow pellet and avoid transferring any cellular debris to the new tubeĪdd 275 µl of ZymoPURE™ Binding Buffer to the cleared lysate from step 7 and Incubate the neutralized lysate on ice for 5 minutes.Ĭentrifuge the neutralized lysate for 5 minutes at 13,000 rpm (~17,900 x g). The sample will turn yellow when the neutralization is complete and a yellowish precipitate will form. Do not vortex! Invert the tube an additional 3-4 times after the sample turnsĬompletely yellow. Do not vortex! Let sit at room temperature for 2-3 minutes.Ĭells are completely lysed when the solution appears clear, purple, and viscousĪdd 250 µl of ice cold ZymoPURE™ P3 (Yellow) and mix thoroughly by inversion. To each pellet, add 250 µl of ZymoPURE™ P1 (Red) to the bacterial cell pellet and resuspend completely by vortexing or pipetting.Īdd 250 µl of ZymoPURE™ P2 (Green) and immediately mix by gently inverting the tube 6-8 times. They are labeled "pPSU1 α" and "pPSU2 α", as the growth strain is DH5alpha.Įach pellet was prepared by spinning 1.5 mL of overnight culture at 8,000 rpm (6,800 x g) for 3 minutes. The following protocol is copied from Zymo Research.įrom the freezer, collect pelleted overnight liquid cultures. coli culture using the ZymoPure Plasmid Miniprep Kit and elute with water. Harvest pPSU1 and pPSU2 DNA from overnight E. How do the plasmids' replication relate to that of pBR322 ( genbank)? How many copies of the plasmid would you roughly expect in each cell? Where do PstI and EcoRV cut within their recognition sequences? What are the distances between the EcoRV (GATATC) sites in each plasmid? What are the distances between the PstI (CTGCAG) sites in each plasmid? What software are you using to read the sequence files? Remember to always initial and label your tubes so you can identify their contents and distinguish them from your classmates' material.ĭownload and review the genbank sequence files for pPSU1 and pPSU2. To help determine the recognition sequence for the restriction enzyme AvaII. George published a 1978 paper that sequenced the plasmid pBR322 They were harnessed in the 1970'sĪs a foundational technology for mapping and recombining DNA across all forms of life. Restriction enzymes are among the natural defense systems of bacteria to protect against viral bacteriophages. Restriction enzymes, which cut DNA at sequence-specific The enzymes we will use for our example lab are 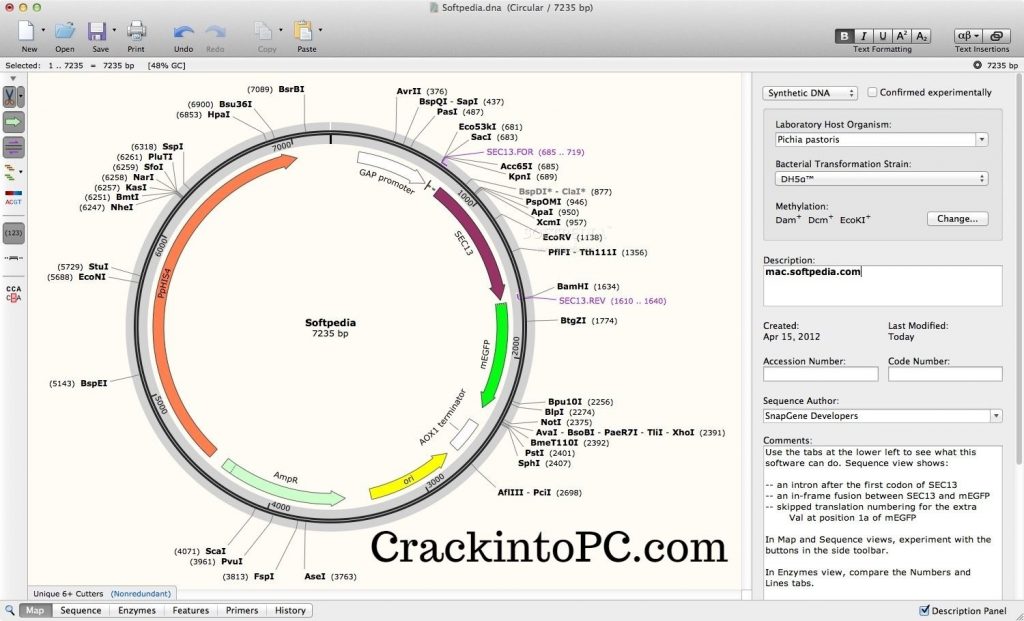
Note, DNA ladders are important references when measuring the size and topology of DNA productsĪfter an enzymatic reaction. How their plasmid designs can produce cheap DNA ladder. In a 2017 paper, Henrici et al demonstrated PPSU2 deposited by the Tan lab at Penn State University. We will be working with Addgene plasmids pPSU1 and Thermocycler, water bath, incubator, or heat blockīenchling, Snapgene, Geneious, ApE, GenBeans, or text editor TAE Buffer (Tris-acetate-EDTA), Agarose, Loading dye,ĭNA stain, 100bp and 1kbp molecular-weight markers (DNA ladders) Hardware, Software, Wetware FunctionĪddgene pPSU1 and pPSU2 bacterial stabs in vials Isolate that DNA from cells, run an enzymatic reaction on it, and show the change you made. Read and annotate the sequence files for a DNA molecule.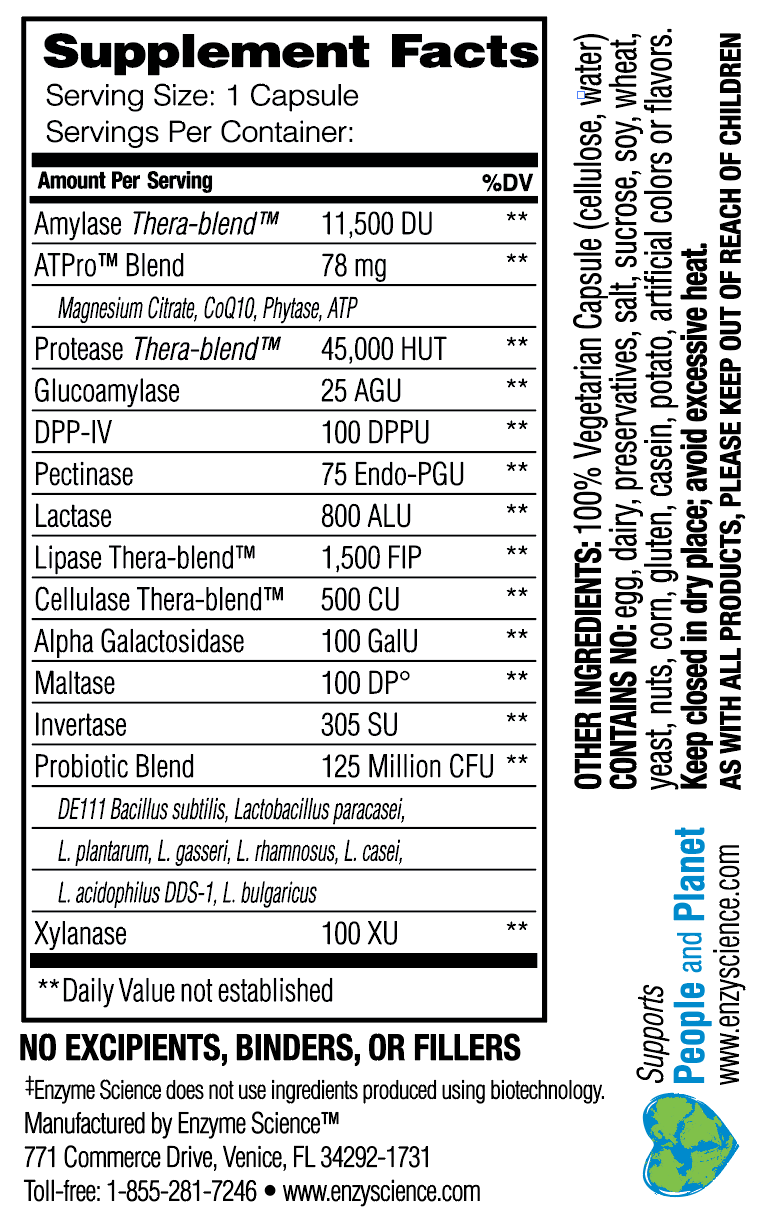 
Set up your student git page and upload your homework from the Principles and Practices class. In your groups, share 2 slides (1 min each) on potential final project ideas. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
